What are Motifs and Domains?
Proteins are classified into a hierarchy of super families, families, and subfamilies according to their relatedness in structure and function [1]. This classification aids in understanding evolutionary history of a protein, as well as determining structure and function of newly discovered proteins. When a protein's amino acid sequence is determined, it may contain signature sequences called motifs. Motifs may confer binding sites or active sites of a protein, or portions of the protein that are modified as it is synthesized. Motifs are often part of a protein domain, part of a gene that is recognized as a unit of a protein. Domains are like pieces of a protein that can be seen in different families but perform the same role due to their amino acid composition.
Conserved Domains of Glyoxalase Family
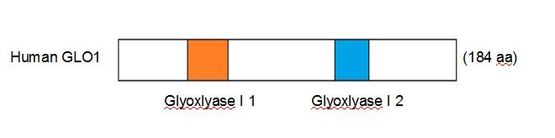
Figure 1. signatures of human GLO1
GLO1 is a member of the cl14632 super family, a group of metalloproteins [2]. Glyoxalase 1 binds Zn(2+) at a Histidine residue located within the second of two domains shown in Fig. 1. These domains are highly conserved throughout all homologs of glyoxalase 1, distinct in their sequence and the distance between them.
Glyoxalase I 1
The first signature of the glyoxalase family is responsible for the enzyme's lactoylglutathione lyase activity. Here the enzyme binds its substrates glutathione and methyglyoxal, catalyzing their conversion to S-D-lactoyl-glutathione (which is further metabolized into D-lactate by glyoxalase II in the glyoxalase system, seen on the protein description tab, Fig. 2).
The sequence spans aa34-55 in the gene in humans. Its consensus sequence is as illustrated [3]:
[HQ]-[IVT]-x-[LIVFY]-x-[IV]-x(4)-{E}-[STA]-x(2)-F-[YM]-x(2,3)-[LMF]-G-[LMF]
The sequence spans aa34-55 in the gene in humans. Its consensus sequence is as illustrated [3]:
[HQ]-[IVT]-x-[LIVFY]-x-[IV]-x(4)-{E}-[STA]-x(2)-F-[YM]-x(2,3)-[LMF]-G-[LMF]
Glyoxalase I 2
The second characterizing domain of glyoxalase 1 is signature 2, responsible for metal ion binding. GLO1 is specific to bind Zn(2+), enabling its catalytic activity.
The sequence spans aa118-134 in the gene in humans. Its consensus sequence is as illustrated [3]:
G-[NTKQ]-x(0,5)-[GA]-[LVFY]-[GH]-H-[IVF]-[CGA]-x-[STAGLE]-x(2)-[DNC]
The sequence spans aa118-134 in the gene in humans. Its consensus sequence is as illustrated [3]:
G-[NTKQ]-x(0,5)-[GA]-[LVFY]-[GH]-H-[IVF]-[CGA]-x-[STAGLE]-x(2)-[DNC]
References
[1] "Introduction to Protein Classification at the EBI." Protein Classification. Retrieve February 28, 2013 from http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification
[2] Marchler-Bauer A et al. (2013), "CDD: conserved domains and protein three-dimensional structure.", Nucleic Acids Res. 41(D1):D384-52.
[3] Sigrist CJA, de Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I. New and continuing developments at PROSITE. Nucleic Acids Res. 2012; doi: 10.1093/nar/gks1067
PubMed: 23161676 [Full text] [PDF version]
[2] Marchler-Bauer A et al. (2013), "CDD: conserved domains and protein three-dimensional structure.", Nucleic Acids Res. 41(D1):D384-52.
[3] Sigrist CJA, de Castro E, Cerutti L, Cuche BA, Hulo N, Bridge A, Bougueleret L, Xenarios I. New and continuing developments at PROSITE. Nucleic Acids Res. 2012; doi: 10.1093/nar/gks1067
PubMed: 23161676 [Full text] [PDF version]
Site created by: Emma Baar
Last updated: 5-14-2013
University of Wisconsin - Madison
Last updated: 5-14-2013
University of Wisconsin - Madison
